metapr2 - a new 18S metabarcode database.
We are very pleased to announce the new metaPR2 metabarcode database.
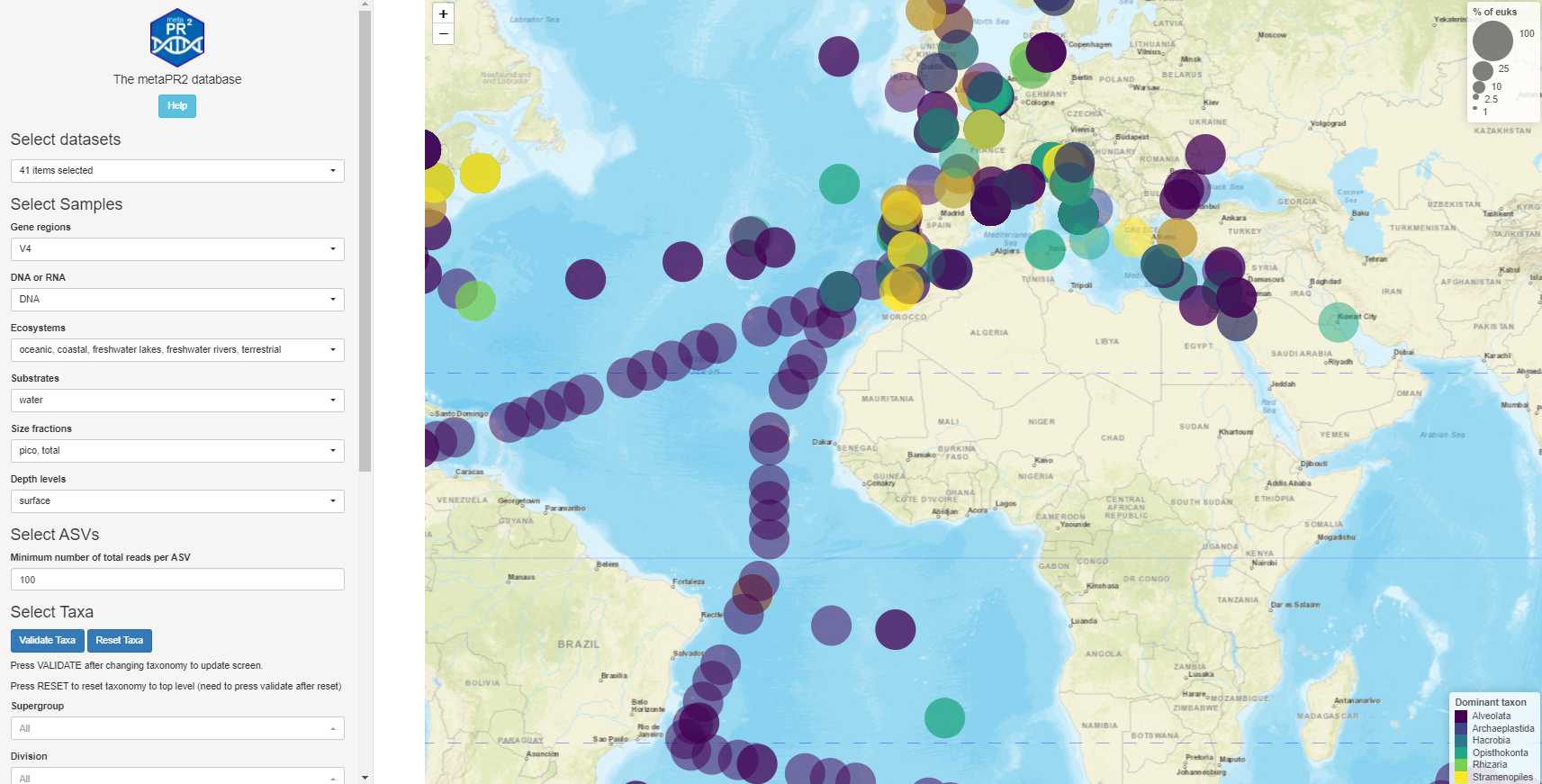
In recent years, metabarcoding has become the method of choice for investigating the composition and assembly of microbial eukaryotic communities. The number of environmental datasets published has increased very rapidly. Although unprocessed sequence files are often publicly available, processed data, in particular clustered sequences, are rarely available in a usable format. Clustered sequences are reported as operational taxonomic units (OTUs) with different similarity levels or more recently as amplicon sequence variants (ASVs). This hampers comparative studies between different environments and datasets, for example examining the biogeographical patterns of specific groups/species, as well analysing the genetic micro-diversity within these groups. We developed the metaPR2 database of processed 18S rRNA metabarcodes that are annotated with the PR2 reference sequence database. Version 1.1 of the database contains 41 datasets corresponding to more than 4,000 samples and 90,000 ASVs. The database, which is accessible through both a web-based interface (https://shiny.metapr2.org) and an R package, should prove very useful to all researchers working on protist diversity in a variety of systems.
Resources
Releated projects
- Ocean Barcode Atlas - Restricted to Tara V9 and Malapsina V4; taxonomy not up to date.
- GlobalFungi - Using the ITS markers and the UNITE reference database.
- MyGOD - The aim of the VDGOB project is to provide interfaces to visualize, explore, analyze and interpret Genomic Observatories data.
Maintainers
- Daniel Vaulot: vaulot@gmail.com
- Adriana Lopes dos Santos: lopesas.ufrj@gmail.com
Citation
Vaulot, D., Sim, C.W.H., Ong, D., Teo, B., Biwer, C., Jamy, M., Lopes dos Santos, A., 2022. metaPR2: a database of eukaryotic 18S rRNA metabarcodes with an emphasis on protists. Molecular Ecology Resources. DOI: 10.1111 / 1755-09