The PR2 databases
Three interconnected 18S rRNA databases

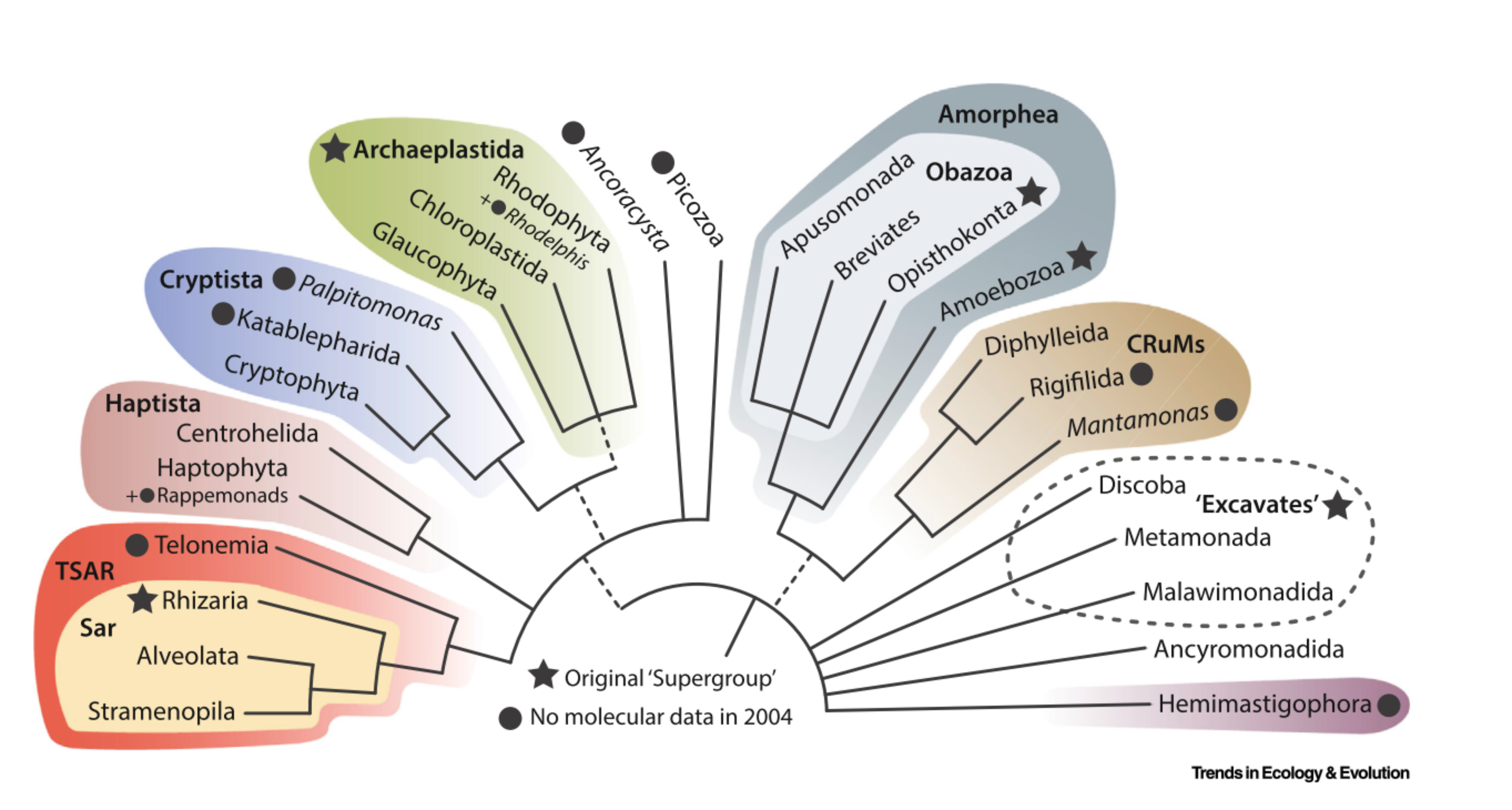
The PR2 (Protist Ribosomal Reference) database ecosystem is a set of three interconnected 18S rRNA databases that are useful in particular for metabarcoding applications.

The PR2 reference sequence database was initiated in 2010 in the frame of the BioMarks project from work that had developed in the previous ten years in the Plankton Group of the Station Biologique of Roscoff. Its aim is to provide a reference database of carefully annotated 18S rRNA sequences using nine unique taxonomic fields (from domain to species). At present it contains over 220,000 sequences. Although it focuses on protists, it also contains sequences from metazoa, fungi and plants as well a limited set of 16S sequences from plastids and bacteria. A number of metadata fields are available for many sequences, including geo-localisation, whether it originates from a culture or a natural sample, host type etc… The annotation of PR2 is performed by experts from each taxonomic groups. One important project in this respect is EukRef which is now part of PR2. EukRef has built bioinformatics pipelines that have been used during three workshops dedicated to specific taxonomic groups. As an example, part of the ciliate annotation originate from the first EukRef workshop. Now the PR2 web interface provides also access to the ROD database that contains full-length eukaryotic ribosomal operons and to . The database is based on the genome assemblies from NCBI, and the operons are extracted from the assemblies. The database currently contains 69,480 operon variants from more than 11,935 genomes.
We have developed an interactive pr2-primers database focusing on the rRNA operon. You can download primer and primer sets. Primer pairs are evaluated against the PR2 sequence database and you can test your own primers and primer sets. To know more.
The metaPR2 database contains processed 18S rRNA metabarcodes that are annotated with the PR2 reference sequence database. Version 3.0 of the database contains 67 datasets corresponding to more than 7,000 samples and 90,000 ASVs. To know more.
Guillou, L., Bachar, D., Audic, S., Bass, D., Berney, C., Bittner, L., Boutte, C. et al. 2013. The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote Small Sub-Unit rRNA sequences with curated taxonomy. Nucleic Acids Res. 41:D597–604.
del Campo J., Kolisko M., Boscaro V., Santoferrara LF., Nenarokov S., Massana R., Guillou L., Simpson A., Berney C., de Vargas C., Brown MW., Keeling PJ., Wegener Parfrey L. 2018. EukRef: Phylogenetic curation of ribosomal RNA to enhance understanding of eukaryotic diversity and distribution. PLOS Biology 16:e2005849. DOI: 10.1371/journal.pbio.2005849.
Vaulot, D., Mahé, F., Bass, D., & Geisen, S. (2021). pr2-primer : An 18S rRNA primer database for protists. Molecular Ecology Resources. DOI: 10.1111⁄1755-0998.13465
Vaulot, D., Sim, C.W.H., Ong, D., Teo, B., Biwer, C., Jamy, M., Lopes dos Santos, A., 2022. metaPR2: a database of eukaryotic 18S rRNA metabarcodes with an emphasis on protists. Molecular Ecology Resources. DOI: 10.1111⁄1755-0998.13674
Krabberød, A.K., Stokke, E., Thoen, E., Skrede, I. & Kauserud, H. 2025. The Ribosomal Operon Database: A Full-Length rDNA Operon Database Derived From Genome Assemblies. Molecular Ecology Resources. 25:e14031.
González-Miguéns, R., Gàlvez-Morante, À., Skamnelou, M., Antó, M., Casacuberta, E., Richter, D.J., Lara, E. et al. 2025. A novel taxonomic database for eukaryotic mitochondrial cytochrome oxidase subunit I gene (eKOI), with a focus on protists diversity. Database (Oxford). 2025:baaf057.
You can contribute to the PR2 database in different ways.
Welcome to the PR2 databases BlueSky account. We will post here important news about the PR2 sequence databases as well as papers using the databases. pr2-database.org
— The PR2 database (@pr2-database.bsky.social) 19 November 2024 at 13:35
[image or embed]
 |
 |
 |
 |
 |
 |